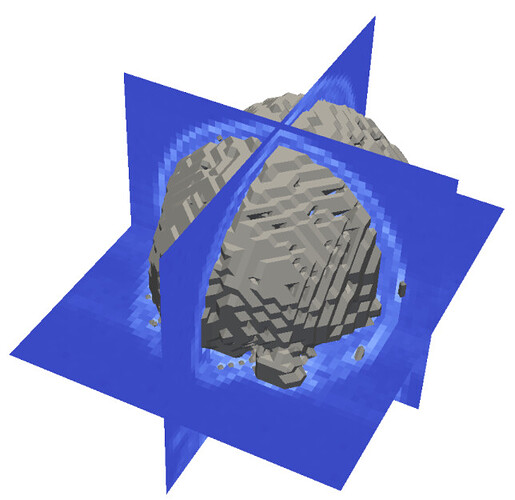
However, I can confirm that there is a difference between the DICOM tags and resulting Mimics project. Unfortunately we don’t have much experience with the NiBabel package and required affine transformation ourselves so I’m afraid we might be very limited in how we can help as we want to avoid giving wrong information. the axial plane image (second column, bottom row) is in the locatin for the coronal image and it needs to be rotated 90 degrees the coronal plane image (second column, top row) is in the location for the sagittal image and it needs to be rotated clockwise by 90 degrees the sagittal plane image (first column, top row of the image quadrant) is in the location for the axial image However, the resulting images (showing here the mask of a deformed vertebral column) are oriented all wrong, when I view it in ITK-snap. Mask_value = mimics_mask.get_voxel_buffer() I obtain the mask using the scripting commands: Nifti patient = Dicom image where the resulting rotation matrix is: To obtain the nifti patient orientation (world), I would do the following: Therefore, i and j are inverted with respect to Dicom and the Dicom patient to Nifti patient transformation matrix is Therefore, the transformation affine matrix is from Dicom to Patient (assuming pixel spacing and slice thickness of 1 to keep it simple) and switching/flipping rows and columns as mentioned in the web page referenced above:įrom what I have read, the Nifti format has a different patient coordinate system than the Dicom, where right is +, anterior is +, and superior is+. This means that the images coordinate system is left+, inferior+ and posterior+. In my images, I get the following info from this tag:įrom my readings, I understand that the patient/world coordinate for Dicom standard is left +, posterior +, superior +. I am using this page as a source of information: Neuroimaging in Python - NiBabel 3.2.1+100.g9fdf5e3e documentationįrom what I understand, the tag 0020, 0037 provides the orientation of the rows and column of the slice of the MRI (in this case) image to transform the image into the patient (dicom) coordinate system. I think it is a really silly mistake but for some reason I cannot figure it out. However, I have a question regarding affine transformations as I am getting something wrong in the process. I am trying to export the masks in mimics to nifti format, via the scripting module. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
